Antigen presenting dendritic cells (DC) represent highly specialized immune cells with a central role in immunity and tolerance induction. DC sense antigens, which are taken-up, processed and presented in the context of MHC molecules to elicit antigen specific T cell responses. Specific DC subsets exist that differ in surface phenotype, function, activation state and anatomical localization, including (i) classical DC type 1 and 2 (cDC1 and cDC2, respectively) in lymphoid and non-lymphoid tissues; (ii) plasmacytoid DC (pDC) in blood that represent the major producers of type 1 interferon and (iii) Langerhans cells (LC), the cutaneous contingent of DC in epidermis.
 |
Langerhans cells of epidermal skin are stained for MHC class II (green) |
The helix-loop-helix transcription factor Id2 represents a determining factor for DC development (Hacker et al., 2003; Zenke and Hieronymus, 2006; Seré et al., 2012). Id2-/- mice lack LC and cDC1. TGFbeta1-/- mice also lack LC and we show that TGFbeta1 acts upstream of Id2 and induces Id2 expression.
We identified two types of LC: short-term LC and long-term LC cells (Seré et al., 2012). Short-term LC develop from Gr-1+ monocytes under inflammatory conditions and are Id2-independent. Long-term LC arise from bone marrow under steady state and depend on Id2. LC reconstitution after inflammation occurs in two waves: an initial fast wave of Gr-1+ monocyte-derived short-term LC, which is followed by a second wave of bone marrow-derived long-term LC.
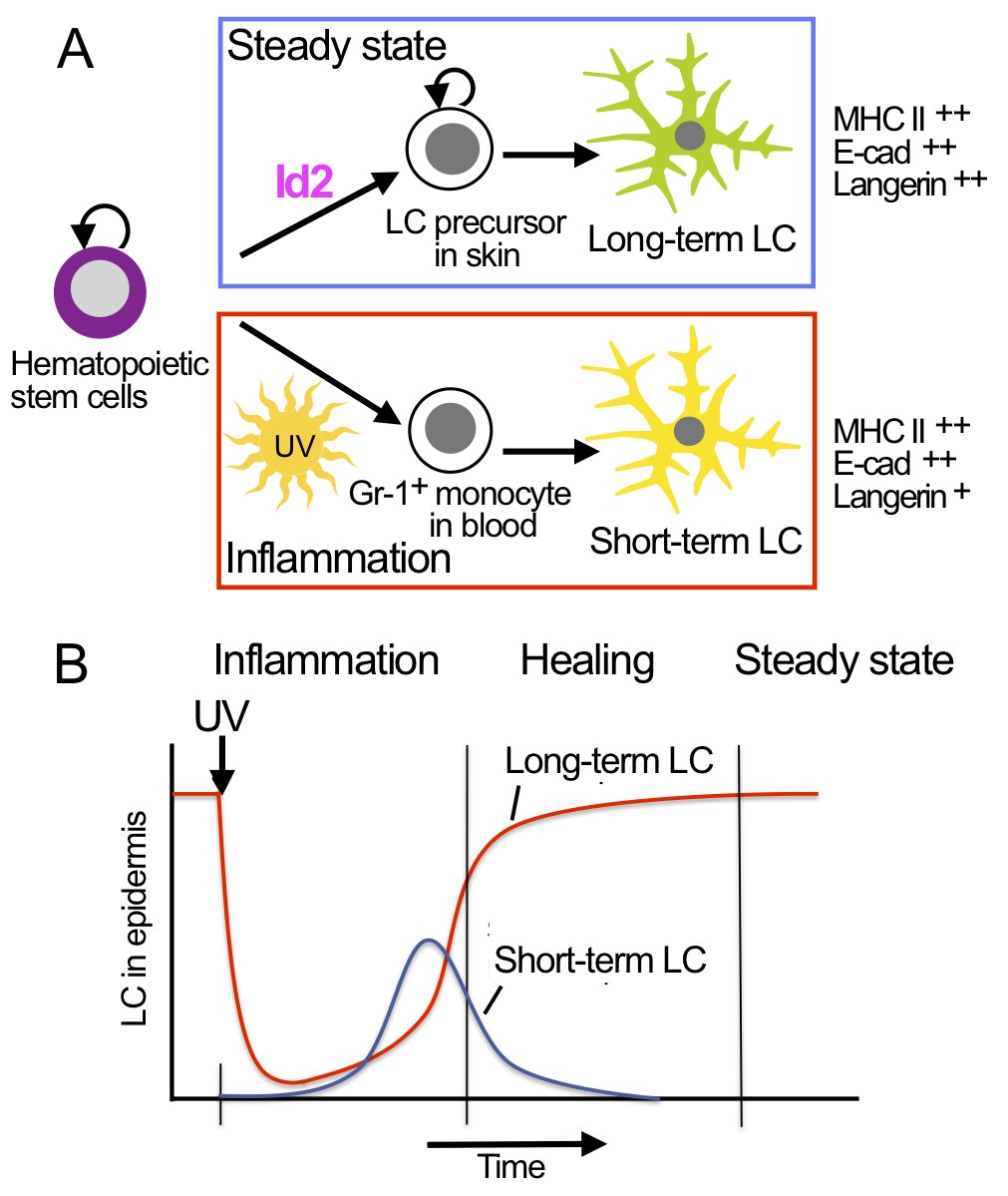 |
Hematopoietic stem cells and LC precursors in skin develop into long-term LC in steady state, which requires the transcription factor Id2 (A, top panel). In inflammation Gr-1+ monocytes develop into short-term LC, which does not require Id2 (A, lower panel). LC development in inflammation occurs in consecutive waves of short-term LC and long-term LC (B). |
Epigenetic and Transcriptional Architecture of Dendritic Cell Development
DC subsets develop from hematopoietic stem cells via Flt3 expressing progenitors through consecutive steps of lineage commitment and differentiation: multipotent progenitors (MPP) are committed to DC restricted common DC progenitors (CDP), which differentiate into the specific DC subsets cDC1 cDC2 and pDC. The laboratory studies gene expression and chromatin architecture of the MPP-CDP-cDC/pDC sequel by employing genome wide approaches with RNA-Seq, ChIP-Seq, ATAC-Seq and HiChIP-seq and a rich toolbox of bioinformatics (Hieronymus et al., 2005; Felker et al., 2010; Chauvistré et al., 2014; Lin et al., 2015; Gusmao et al., 2016; Li et al., 2019; in collaboration with Ivan Costa, Institute for Computational Genomics, RWTH Aachen University, Aachen, Germany).
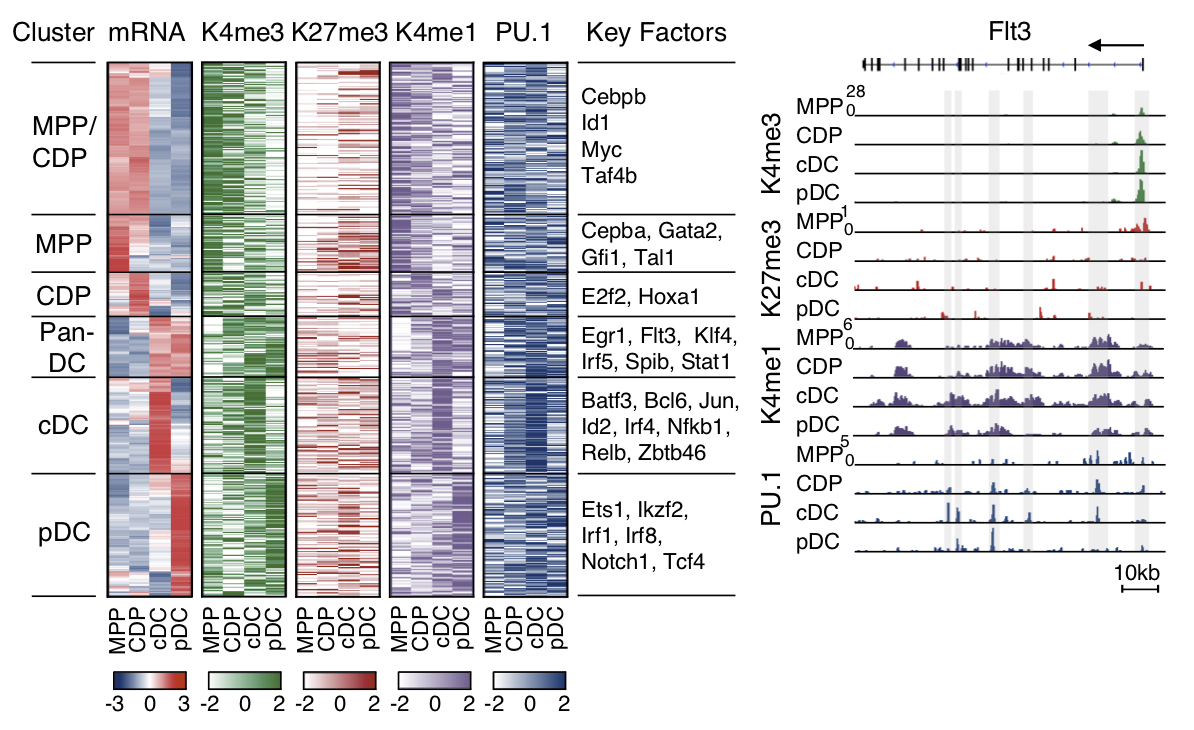 |
Gene expression (mRNA), H3K4me1, H3K4me3, H3K27me3 and PU.1 occupancy of the sequel MPP-CDP-cDC/pDC (left). Histone and PU.1 profile of Flt3 gene (right; Lin et al., 2015). |
Specific H3K4me1, H3K4me3 and H3K27me3 marks in CDP reveal a DC-primed epigenetic signature, which is maintained and reinforced during DC differentiation. We describe the circuitry of transcription factors, including Irf4, Irf8, Tcf4, Spib and Stats, which drives the sequel MPP-CDP-cDC/pDC. The circuitry also includes positive feedback loops inferred for individual or multiple factors that stabilize the distinct stages of DC development.
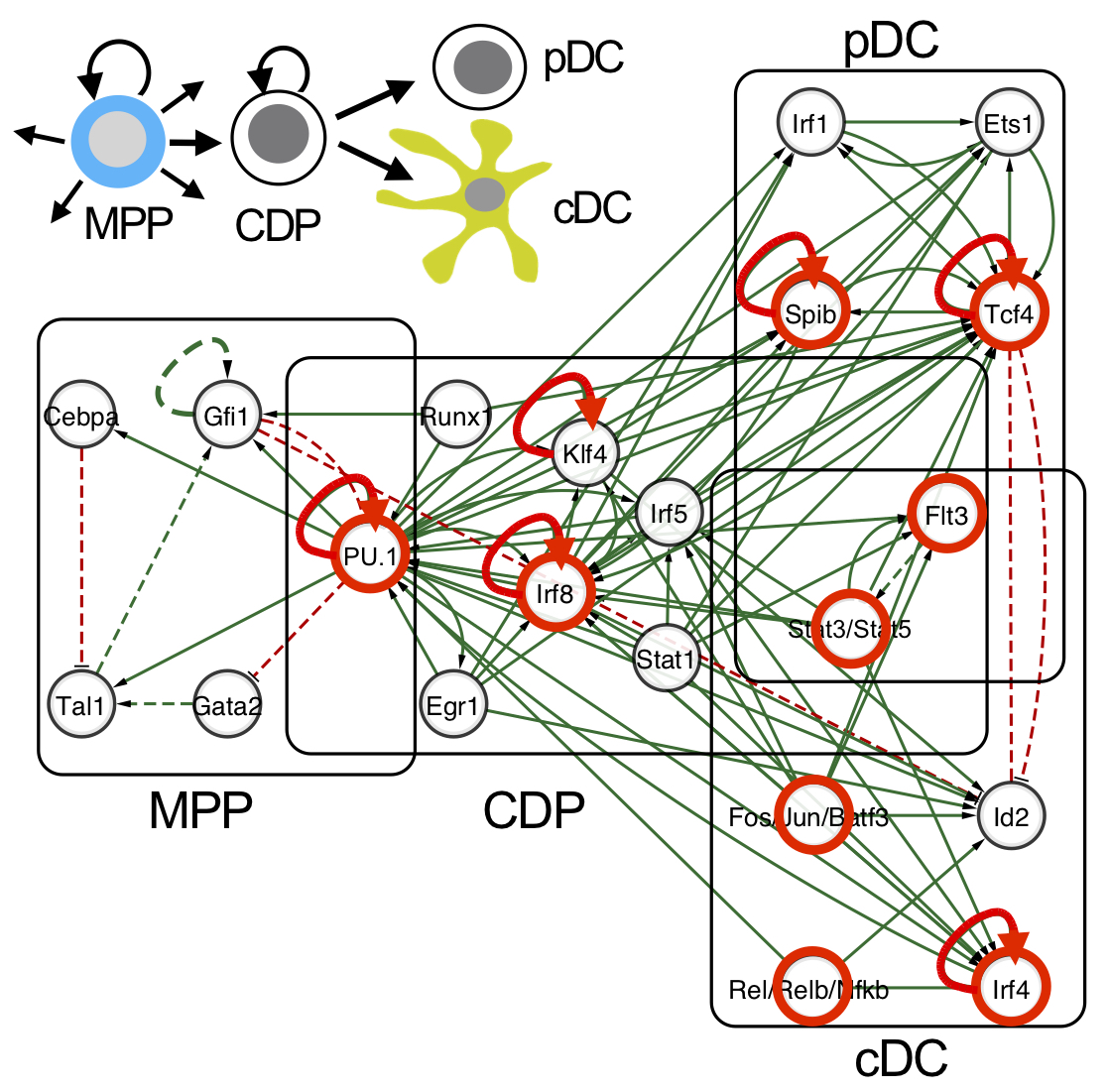 |
Network illustration of the integrated DC regulatory circuitry of the sequel MPP-CDP-cDC/pDC (Lin et al., 2015). Transcription factors that impact on DC development in gene knockout studies are indicated in red. Feedback loops that stabilize the distinct DC stages in the network are depicted (red arrows). |
Data are available by the customized UCSC genome browser track data hub http://www.molcell.rwth-aachen.de/dc/.
Selected publications:
| A lncRNA identifies IRF8 enhancer element in negative feedback control of dendritic cell differentiation.. Xu, H., Li, Z., Kuo, C.-C. Götz, K., Look, T., Toledo, M. A. S., Seré, K., Costa, I. G. and Zenke, M. (2023). |
||
| eLife 12, e83342. | Abstract | Full | |
| Intrathymic dendritic cell precursors promote human T-lineage specification via IRF8- driven transmembrane TNF. Liang, K. L., Roels, J., Lavaert, M., Putteman, T., Boehme, L., Tilleman, L., Velghe, I., Pegoretti, V., van de Walle, I., Sontag, S., Vandewalle, J., Vandekerckhove, B., Leclercq, G., van Vlierberghe, P., Libert, C., van Nieuwerburgh, F., Fischer, R., Kontermann, R. E., Pfizenmaier, K., Doody, G., Zenke, M., and Taghon, T. (2022). |
||
| Nat. Immunol. 24, 474-486. | Abstract | Full | |
| Guidelines for mouse and human dendritic cell generation. Lutz, M. B., Ali, S. Audiger, C., Autenrieth, S. E., Berod, L., Bigley, V., Cyran, L., Dalod, M., Dörrie, J., Dudziak, D., Flórez-Grau, G., Giusiano, L., Godoy, G. J., Heuer, M., Krug, A. B., Lehmann, C. H. K., Mayer, C. T., Naik, S. H., Scheu, S., Schreibelt, G., Segura, E., Seré, K., Sparwasser, T., Tel, J., Xu, H. and Zenke, M. (2022). |
||
| Eur. J. Immunol., doi: 10.1002/eji.202249816. Online ahead of print. | Abstract | Full | |
| Dendritic cells generated from induced pluripotent stem cells and by direct reprogramming of somatic cells. Flosdorf, N., and Zenke, M. (2022). |
||
| Eur. J. Immunol. 52, 1880-1888. | Abstract | Full | |
| CRISPR/Cas9 editing in conditionally immortalized HoxB8 cells for studying gene regulation in mouse dendritic cells. Xu, H., Look, T., Prithiviraj, S., Lennartz, D., Delgado Cáceres, M., Götz, K., Wanek, P., Häcker, H., Kramann, R., Seré, K., and Zenke, M. (2021). |
||
| Eur. J. Immunol. 52, 1859-1862. | ||
| Chromatin-accessibility estimation from single-cell ATAC data with scOpen. Li, Z., Kuppe, C., Ziegler, S., Cheng, M., Kabgani, N., Menzel, S., Zenke, M., Kramann, R., and Ivan G. Costa, I. G. (2021). |
||
| Nature Communications 12, 6386. | ||
| Human DC3 antigen presenting dendritic cells from induced pluripotent stem cells. Satoh, T., Toledo, M. A. S., Boehnke, J., Olschok, K., Küstermann, C., Sontag, S., Seré, K., Koschmieder, S., Brümmendorf, T. H., Chatain, N., Tagawa, Y.-I., and Zenke, M. (2021). |
||
| Front. Cell Dev. Biol. 9, 667304. | ||
| Epigenetic aspects of DC development and differentiation. Chauvistré, H., and Seré, K. (2020). |
||
| Mol. Immunol. 128,116-124. | ||
| Identification of transcription factor binding sites using ATAC-seq. Li, Z., Schulz, M. H., Look, T., Begemann, M., Zenke, M., and Ivan G. Costa, I. G. (2019). |
||
| Genome Biol. 20, 45. | Abstract | Full | |
| The impact of the epithelial–mesenchymal transition regulator hepatocyte growth factor receptor/Met on skin immunity by modulating Langerhans cell migration. Sagi, Z., and Hieronymus, T. (2018). |
||
| Front. Immunol. 16, 517. | Abstract | Full | |
| Analysis of computational footprinting methods for DNase sequencing experiments. Gusmao, E. G., Allhoff, M., Zenke, M., and Costa, I. G. (2016). |
||
| Nature Methods 13, 303-309 (see also Editorial Nature Methods 13, 185) | Abstract | Full | |
| Epigenetic program and transcription factor circuitry of dendritic cell development. Lin, Q., Chauvistré, H., Costa, I. G., Gusmão, E. G., Mitzka, S., Haenzelmann, S., Baying, B., Klisch, T., Moriggl, R., Hennuy, B., Smeets, H., Hoffmann, K., Benes, V., Seré, K., and Zenke, M. (2015). |
||
| Nucl. Acid Res. 43, 9680-9693. | Abstract | Full | |
| The clash of Langerhans cell homeostasis: Should I stay or should I go? Hieronymus, T., Zenke, M., Baek, J. H., Seré K. (2014). |
||
| Semin. Cell Dev. Biol., S1084-9521. | Abstract | Full | |
| Dendritic cell development requires histone deacetylase activity. Chauvistré, H., Küstermann, C., Rehage, N., Klisch, T., Mitzka, S., Felker, P., Rose-John, S., Zenke, M., and Seré, K. (2014). |
||
| Eur. J. Immunol. 44, 2478-2488. | ||
| Two distinct types of Langerhans cells populate the skin during steady state and inflammation. Seré, K., Baek, J. H., Ober-Blöbaum, J., Müller-Newen, G., Tacke, F., Yokota, Y., Zenke, M., and Hieronymus, T. (2012). |
||
| Immunity 37, 905-916. | ||
| (see also Preview by Romani, N., Tripp, C. H. and Stoitzner, P. (2012). Langerhans cells come in waves. Immunity 37, 766-768). | ||
| The HGF receptor/met tyrosine kinase is a key regulator of dendritic cell migration in skin immunity. Baek, J. H., Birchmeier, C., Zenke, M., and Hieronymus, T. (2012). |
||
| J. Immunol. 189, 1699-1707. | ||
| TGF-ß1 accelerates dendritic cell differentiation from common dendritic cell progenitors and directs subset specification toward conventional dendritic cells. Felker, P., Seré, K., Lin, Q., Becker, C., Hristov, M., Hieronymus, T., and Zenke, M. (2010). |
||
| J. Immunol. 185, 5326-5335. | ||
| Transforming growth factor beta1 up-regulates interferon regulatory factor 8 during dendritic cell development. Ju, X.-S., Ruau, D., Jäntti, P., Sere, K., Becker, C., Wiercinska, E., Bartz, C., Erdmann, B., Dooley, S., and Zenke, M. (2007). |
||
| Eur. J. Immunol. 37, 1174-1183. | ||
| Towards an understanding of the transcription factor network of dendritic cell development. Zenke, M., and Hieronymus, T. (2006). |
||
| Trends Immunol. 27, 140-145. | ||
| Progressive and controlled development of mouse dendritic cells from Flt3+CD11b+ progenitors in vitro. Hieronymus, T., Gust, T. C., Kirsch, R. D., Jorgas, T., Blendinger, G., Goncharenko, M., Supplitt, K., Rose-John, S., Muller, A. M., and Zenke, M. (2005). |
||
| J. Immunol. 174, 2552-2562. | ||
| Transcriptional profiling identifies Id2 function in dendritic cell development. Hacker. C., Kirsch, R. D., Ju, X.-S., Hieronymus, T., Gust, T. C., Kuhl, C., Jorgas, T., Kurz, S. M., Rose-John, S., Yokota, Y., and Zenke, M. (2003). |
||
| Nat. Immunol. 4, 380-386. | ||
| MHC class II presentation of endogenously expressed antigens by transfected dendritic cells. Diebold, S. S., Cotten, M., Koch, N., and Zenke, M. (2001). |
||
| Gene Ther. 8, 487-493. |
Abstract| Full
|
|
| Dendritic cell progenitor is transformed by a conditional v-Rel estrogen receptor fusion protein v-RelER. Boehmelt, G., Madruga, J., Dorfler, P., Briegel, K., Schwarz, H., Enrietto, P. J., and Zenke, M. (1995). |
||
| Cell 80, 341-352. |
Abstract | Full
|
|